[R7I7]
Humans can rapidly visually recognise the actions people around them are engaged in and this ability is important for successful behaviour and social interaction. Isik et al. presented human subjects with 2-second video clips of humans performing actions while measuring brain activity with MEG. The clips comprised 5 actions (walk, run, jump, eat, drink) performed by each of five different actors and video-recorded from each of five different views (only frontal and profile used in MEG). Results show that action can be decoded from MEG signals arising about 200 ms after the onset of the video, with decoding accuracy peaking after about 500 ms and then decaying while the stimulus is still on, with a rebound after stimulus offset. Moreover, decoders generalise across actors and views. The authors conclude that the brain rapidly computes a representation that is invariant to view and actor.
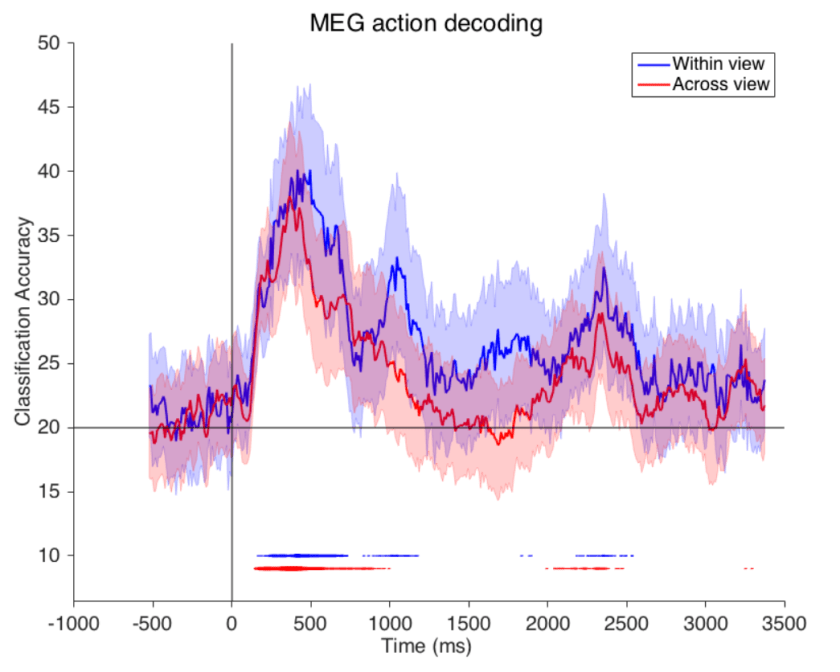
Figure from the paper. Legend from the paper with my modifications in brackets: [Accuracy of action decoding (%) from MEG data as a function of time after video onset]. We can decode [which of five actions was being performed in the video clip] by training and testing on the same view (‘within-view’ condition), or, to test viewpoint invariance, training on one view (0 degrees [frontal, I think, but this should be clarified] or 90 degrees [profile]) and testing on the second view (‘across view’ condition). Results are each [sic] from the average of eight different subjects. Error bars represent standard deviation [across what?]. Horizontal line indicates chance decoding accuracy. […] Lines at the bottom of plot indicate significance with p<0.01 permutation test, with the thickness of the line indicating [for how many of the 8 subjects decoding was significant]. [Note the significant offset response after the 2-s video (whose duration should be indicated by a stimulus bar).]
The rapid view-invariant action decoding is really quite amazing. It would be good to see more detailed analyses to assess the nature of the signals enabling this decoding feat. Of course, 200 ms already allows for recurrent computations and the decodability peak is at 500 ms, so this is not strong evidence for a pure feedforward account.
The generalisation across actors is less surprising. This was a very controlled data set. Despite some variation in the appearance of the actors, it seems plausible that there would be some clustering of the vectors of pixels across space and time (or of features of a low-level visual representation) corresponding to different actors performing the same action seen from the same angle.
In separate experiments, the authors used static single frames taken from the videos and dynamic point-light figures as stimuli. These reduced form-only and motion-only stimuli were associated with diminished separation of actions in the human brain and in model representations, and with diminished human action recognition, suggesting that form and motion information are both essential to action recognition.
I’m wondering about the role of task-related priors. Subjects were performing an action recognition task on this controlled set of brief clips during MEG while freely viewing the clips (though this is not currently clearly stated). This task is likely to give rise to strong prior expectations about the stimulus (0 deg or 90 deg, one of five actions, known scale and positions of key features for action discrimination). Primed to attend to particular diagnostic features and to fixate in certain positions, the brain will configure itself for rapid dynamic discrimination among the five possible actions. The authors present a group-average analysis of eye movements, suggesting that these do not provide as much information about the actions as the MEG signal. However, the low-dimensional nature of the task space is in contrast to natural conditions, where a wider variety of actions can be observed and view, actor, size, and background vary more. The precise prior expectations might contribute to the rapid discriminability of the actions in the brain signals.
The authors model the results in the framework of feedforward processing in a Hubel-and-Wiesel/Poggio-style model that alternates convolution and max-pooling to simulate responses resembling simple and complex cells, respectively. This model is extended here to process video using spatiotemporal filter templates. The first layer uses Gabor filters, higher layers use templates in the first layer matching video clips in the stimulus set. The authors argue that this model supports invariant decoding and largely accounts for the MEG results.
Like the subjects, the model is set up to process the restricted stimulus space. The internal features of the model were constructed using representational fragments from samples from the same parametric space of videos. The exact videos used to test the models were not used for constructing the feature set. However, if I understood correctly, videos from the same restricted space (5 actions, 5 actors, 5 views) were used. Whether the model can be taken to explain (at a high level of abstraction) the computations performed by the brain depends critically on the degree to which the model is not specifically constructed for the (necessarily very limited) 5-action controlled stimulus space used in the study.
As the authors note, humans still outperform computer vision models at action recognition. How does the authors’ own model perform on less controlled action videos? If it the model cannot perform the task on real-world sensory input, can we be confident that it captures the way that the human brain performs the task? This is a concern in many studies and not trivial to address. However, the interpretation of the results should engage this issue.
Strengths
- Controlled stimulus set: The set of video stimuli (5 actions x 5 actors x 5 views x 26 clips = 3250 2-sec clips) is impressive. Assembling this set is an important contribution to the field. The set is condition-rich (compared to typical stimulus sets used in cognitive neuroscience studies) and seems to strike a good balance between control and realism. This set could be a driver of progress if it were to be used in multiple modelling and empirical studies.
- Combination of brain-activity measurements and a simple computational model, which provides a useful starting point for modelling the recognition of dynamic actions, as it is minimal and standard in many respects: a feedforward model in the HMAX framework, extended from spatial to spatiotemporal filters.
Weaknesses
- Controlled stimulus set: The set of video stimuli is very restricted compared to real-world action recognition. For the brain data, this means that subjects might have rapidly formed priors about the stimuli, enabling them to configure their visual systems (e.g. attentional templates, fixation targets) for more rapid recognition of the 5 actions than is possible in real-world settings. This limitation is shared with many studies in our field and difficult to overcome without giving up control (which is a strength, see above). I therefore suggest addressing this problem in the discussion.
- The model uses features based on spatiotemporal patterns sampled from the same restricted stimulus space. Although non-identical clips were used, the videos underlying the representational space appear to share a lot with the experimental stimuli (same 5 actions, same 5 views, same background?, same actors?). I would therefore not expect this model to work well on arbitrary real-world action video clips. This is in contrast to recent studies using deep convolutional neural nets (e.g. Khaligh-Razavi & Kriegeskorte 2014), where the models were trained without any information about the (necessarily restricted) brain-experimental stimulus set and can perform recognition under real-world conditions.
- Only one model (in two variants) is tested. In order to learn about computational mechanism, it would be good to test more models.
- MEG data were acquired during viewing of only 50 of the clips (5 actions x 5 actors x 2 views).
- Missing inferential analyses: While the authors employ inferential analyses in single subjects and report number of significant subjects, few hypotheses of interest are formally statistically tested. The effects interpreted appear quite strong, so the results described above appear solid nevertheless (interpretational caveats notwithstanding).
Overall evaluation
This is an ambitious study describing results of a well-designed experiment using a stimulus set that is a major step forward. The results are an interesting and substantial contribution to the literature. However, the analyses could be much richer than they currently are and the interpretation of the results is not straightforward. Stimulus-set-induced priors may have affected both the neural processing measured and the model (which used templates from stimuli within the controlled video set). Results should be interpreted more cautiously in this context.
Although feedforward processing is an important part of the story, it is not the whole story. Recurrent signal flow is ubiquitous in the brain and essential to brain function. In engineering, similarly, recurrent neural networks are beginning to dominate spatiotemporal processing challenges such as speech and video recognition. The fact that the MEG data are presented as time courses, revealing a rich temporal structure, and the model analyses are bar graphs illustrates the key limitation of the model.
It would be great to extend the analyses to reveal a richer picture of the temporal dynamics. This should include an analysis of the extent to which each model layer can explain the representational geometry at each latency from stimulus onset.
Future directions
In revision or future studies, this line of work could be extended in a number of ways:
- Use multiple models that can handle real-world action videos. The authors’ controlled video set is extremely useful for testing human and model representations, and for comparing humans to models. However, to be able to draw strong conclusions, the models, like the humans, would have to be trained to recognise human actions under real-world conditions (unrestricted natural video). In addition, it would be good to compare the biological representational dynamics to both feedforward and recurrent computational models.
- To overcome the problem of stimulus-set related priors, which make it difficult to compare representational dynamics measured for restricted stimulus sets to real-world recognition in biological brains, one could present a large set of stimuli without ever presenting a stimulus twice to the same subject. Would the action category still be decodable at 150 ms with generalisation across views? Would a feedforward computer vision model trained on real-world action videos be able to predict the representational dynamics?
- The MEG analyses could use source reconstruction to enable separate analyses of the representational dynamics in different brain regions.
- It would be useful to have MEG data for the full stimulus set of 5 actions x 5 actors x 5 viewpoints = 125 conditions. The representational geometries could be analysed in detail to reveal which particular action pairs become discriminable when with what level of invariance.
Particular suggestions for improvements of this paper
(1) Present more detailed results
It would be good to see results separately for each pair of actions and each direction of crossdecoding (0 deg training -> 90 deg testing, and 90 deg training -> 0 deg testing). Regarding the former, eating and drinking involve very similar body postures and motions. Is this reflected in the discriminability of these actions?
Regarding, the decoding generalisation across views, you state:
“We decoded by training only on one view (0 degrees or 90 degrees), and testing on a second view (0 degrees or 90 degrees).”
Was the training set exclusively composed on 0 degree (frontal?) and the test set exclusively of 90 degree (side view?), and vice versa? In case the test set contained instances of both views (though of course, not for the same actor and action), results are more difficult to interpret.
(2) Discuss the caveats to the current interpretation of the results
Discuss the question whether priors resulting from subjects understanding of the restricted stimulus set might have affected the processing of the stimuli. Consider the involvement of recurrent computations within 200 ms and discuss the continuing rise of decodability until 500 ms. Discuss the possibility that the model will not generalise to action recognition in the wild.
(3) Test several control models
Can Gabor, HMAX, and deep convolutional neural net models support similarly invariant action decoding? These models are relatively easy to test, so I think it’s worth considering this for revision. Computer vision models trained on dynamic action recognition could be left to future studies.
(4) Test models by comparison of its representations with the brain representations
The computational model is currently only compared to the data at the very abstract level of decoding accuracy. Can the model predict the representations and representational dynamics in detail? It might be difficult to use the model to predict the measured channels. This would require the fitting of a linear model predicting the measured channels from the model units and the MEG data (acquired for only 5 actions x 5 actors x 2 views = 50 conditions) might be insufficient. However, representational dynamics could be investigated in the framework of representational similarity analysis (50 x 50 representational dissimilarity matrices) following Carlson et al. (2013) and Cichy et al. (2014). Note that this approach does not require fitting a prediction model and so appears applicable here. Either approach would reveal the dynamic prediction of the feedforward model (given dynamic inputs) and where its prediction diverges from the more complex and recurrent processes in the brain. This would promise to give us a richer and less purely confirmatory picture of the data and might show the merits and limitations of a feedforward account.
(5) Perform temporal cross-decoding
Temporal crossdecoding (Carlson et al. 2013, Cichy et al. 2014) could be used to more richly characterise the representational dynamics. This would reveal whether representations stabilise in certain time windows, or keep tumbling through the representational space even as stimuli are continuously decodable.
(6) Improve the inferential analyses
I don’t really understand the inference procedure in detail from the description in the methods section.
“We recorded the peak decoding accuracy for each time bin,…”
What is the peak decoding accuracy for each time bin? Is this the maximum accuracy across subjects for each time bin?
“…and used the null distribution of peak accuracies to select a threshold where decoding results performing above all points in the null distribution for the corresponding time point were deemed significant with P < 0.01 (1/100).”
I’m confused after reading this, because I don’t understand what is meant by “peak”.
The inference procedure for the decoding-accuracy time courses seems to lack formal multiple-testing correction across time points. Given enough subjects, inference could be performed with subject as a random effect. Alternatively, fixed-effects inference could be performed by permutation, averaging across subjects. Multiple testing across latencies should be formally corrected for. A simple way to do this is to relabel the experimental events once, compute an entire decoding time course, and record the peak decoding accuracy across time (or if this is what was done, it should be clearly described). Through repeated permutations, a null distribution of peak accuracies can be constructed and a threshold selected that is exceeded anywhere under H0 with only 5% probability, thus controlling the familywise error rate at 5%. This threshold could be shown as a line or as the upper edge of a transparent rectangle that partially obscures the insignificant part of the curve.
For each inferential analysis, please describe exactly what the null hypothesis was, what event-labels are exchangeable under this null hypothesis, and how the null distribution was computed. Also, explain how the permutation test interacted with the crossvalidation procedure. The crossvalidation should ideally generalise to new stimuli and label permutation be wrapped around this entire procedure.
“Decoding analysis was performed using cross validation, where the classifier was trained on a randomly selected subset of 80% of data for each stimulus and tested on the held out 20%, to assess the classifier’s decoding accuracy.”
Does this apply only to the within-view decoding? In the critical decoding analysis with generalisation across views, it cannot have been 20% of the data in the held-out set, since 0-deg views were used for training and 90-deg views for testing (and vice versa). If only 50% of the data were used for training there, why didn’t performance suffer given the smaller training set compared to the within-view decoding?
It would also be good to have estimates and inferential comparisons of the onset and peak latencies of the decoding time courses. Inference could be performed on a set of single-subject latency differences between two conditions modelling subject as a random effect.
(7) Qualify claims about biological fidelity of the model
The model is not really “designed to closely mimic the biology of the visual system”, rather its architecture is inspired by some of the features of the feedforward component of the visual hierarchy, such as local receptive fields of increasing size across a hierarchy of representations.
(8) Open stimuli and data
This work would be especially useful to the community if the video stimuli and the MEG data were made openly available. To fully interpret the findings, it would also be good to be able to view the movie clips online.
(9) Further clarify the title
The title “Fast, invariant representation for human action in the visual system” is somewhat unclear. What is meant are representations of perceived human actions, not representations for action. “Fast, invariant representation of visually perceived human actions” would be better, for example.
(10) Clarify what stimuli MEG data were acquired for
The abstract states “We use magnetoencephalography (MEG) decoding and a computational model to study action recognition from a novel dataset of well-controlled, naturalistic videos of five actions (run, walk, jump, eat, drink) performed by five actors at five viewpoints.” This suggests that MEG data were acquired for all these conditions. The methods section clarifies that MEG data were only recorded for 50 conditions (5 actions x 5 actors x 2 views). Here and in the legend of Fig. 1, it would be better to use the term “stimulus set” in place of “data set”.
(11) Clarify whether subjects were fixating or free viewing
Were subjects free viewing or fixating? This should be explicitly stated and the choice motivated in either case.
(12) Make figures more accessible
The figures are not optimal. Every panel showing decoding results should be clearly labelled to state what variables the crossvalidation procedure tested for generalisations across. For example, a label (in the figure itself!) could be: “decoding brain representations of actions with invariance to actor and view”. The reader shouldn’t have to search in the legend to find this essential information. Also every figure should have a stimulus bar depicting the period of stimulus presence. This is important especially to assess stimulus-offset-related effects, which appear to be present and significant.
Fig. 3 is great. I think it would be clearer to replace “space-time dot product” with “space-time convolution”.
(13) Clarify what the error bars represent
“Error bars represent standard deviation.”
Is this the standard deviation across the 8 subjects? Is it really the standard deviation or the standard error?
(14) Clarify what we learn from the comparison between the structured and the unstructured model
For the unstructured model, won’t the machine learning classifier learn to select random combinations that tend to pool across different views of one action? This would render the resulting system computationally similar.