[I8R7]
Baldassano, Chen, Zadbood, Pillow, Hasson & Norman (pp2016) investigated brain representations of event sequences with fMRI. The paper argues in favour of an intriguing and comprehensive account of the representation of event sequences in the brain as we experience them, their storage in episodic memory, and their later recall.
The overall story is quite amazing and goes like this: Event sequences are represented at multiple time scales across brain regions during experience. The brain somehow parses the continuous stream of experience into discrete pieces, called events. This temporal segmentation occurs at multiple temporal scales, corresponding perhaps to a tree of higher-level (longer) events and subevents. Whether the lower-level events precisely subdivide higher-level events (rendering the multiscale structure a tree) is an open question, but at least different regions represent event structure at different scales. Each brain region has its particular time scale and represents an event as a spatial pattern of activity. The encoding in episodic memory does not occur continuously, but rather in bursts following the event transitions at one of the longer time scales. During recall from memory, the event representations are reinstated, initially in the higher-level regions, from which the more detailed temporal structure may come to be associated in the lower-level regions. Event representations can arise from perceptual experience (a movie here), recall (telling the story), or from listening to a narration. If the event sequence of a narration is familiar, memory recall can help reinstate representations upcoming in the narration in advance.
There’s previous evidence for event segmentation (Zacks et al. 2007) and multi-time-scale representation (from regional-mean activation to movies that are temporally scrambled at different temporal scales; Hasson et al. 2008; see also Hasson et al. 2015) and for increased hippocampal activity at event boundaries (Ben-Yakov et al. 2013). However, the present study investigates pattern representations and introduces a novel model for discovering the inherent sequence of event representations in regional multivariate fMRI pattern time courses.
The model assumes that a region represents each event k = 1..K as a static spatial pattern mk of activity that lasts for the duration of the event and is followed by a different static pattern mk+1 representing the next event. This idea is formalised in a Hidden Markov Model with K hidden states arranged in sequence with transitions (to the next time point) leading either to the same state (remain) or to the next state (switch). Each state k is associated with a regional activity pattern mk, which remains static for the duration of the state (the event). The number of events for a given region’s representation of, say, 50 minutes’ experience of a movie is chosen so as to maximise within-event minus between-event pattern correlation on a held-out subject.
It’s a highly inspired paper and a fun read. Many of the analyses are compelling. The authors argue for such a comprehensive set of claims that it’s a tall order for any single paper to fully substantiate all of them. My feeling is that the authors are definitely onto something. However, as usual there may be alternative explanations for some of the results and I am left with many questions.
Strengths
- The paper is very ambitious, both in terms brain theory and in terms of analysis methodology.
- The Hidden Markov Model of event sequence representation is well motivated, original, and exciting. I think this has great potential for future studies.
- The overall account of multi-time-scale event representation, episodic memory encoding, and recall is plausible and fascinating.
Weaknesses
- Incomplete description and validation of the new method: The Hidden Markov Model is great and quite well described. However, the paper covers a lot of ground, both in terms of the different data sets, the range of phenomena tackled (experience, memory, recall, multimodal representation, memory-based prediction), the brain regions analysed (many regions across the entire brain), and the methodology (novel complex method). This is impressive, but it also means that there is not much space to fully explain everything. As a result there are several important aspects of the analysis that I am not confident I fully understood. It would be good to describe the new method in a separate paper where there is enough space to validate and discuss it in detail. In addition, the present paper needs a methods figure and a more step-by-step description to explain the pattern analyses.
- The content and spatial grain of the event representations is unclear. The analyses focus on the sequence of events and the degree to which the measured pattern is more similar within than between inferred event boundaries. Although this is a good idea, I would have more confidence in the claims if the content of the representations was explicitly investigated (e.g. representational patterns that recur during the movie experience could represent recurring elements of the scenes).
- Not all claims are fully justified. The paper claims that events are represented by static patterns, but this is a model assumption, not something demonstrated with the data. It’s also claimed that event boundaries trigger storage in long-term memory, but hippocampal activity appears to rise before event boundaries (with the hemodynamic peak slightly after the boundaries). The paper could even more clearly explain exactly what previous studies showed, what was assumed in the model (e.g. static spatial activity patterns representing the current event) and what was discovered from the data (event sequence in each region).
Particular points the authors may wish to address in revision
(1) Do the analyses reflect fine-grained pattern representations?
The description of exactly how evidence is related between subjects is not entirely clear. However, several statements suggest that the analysis assumes that representational patterns are aligned across subjects, such that they can be directly compared and averaged across subjects. The MNI-based intersubject correspondency is going to be very imprecise. I would expect that the assumption of intersubject spatial correspondence lowers the de facto resolution from 3 mm to about a centimetre. The searchlight was a very big (7 voxels = 2.1cm)3 cube, so perhaps still contained some coarse-scale pattern information.
However, even if there is evidence for some degree of intersubject spatial correspondence (as the authors say results in Chen et al. 2016 suggest), I think it would be preferable to perform the analyses in a way that is sensitive also to fine-grained pattern information that does not align across subjects in MNI space. To this end patterns could be appended, instead of averaged, across subjects along the spatial (i.e. voxel) dimension, or higher-level statistics, such as time-by-time pattern dissimilarities, could averaged across subjects.
If the analyses really rely on MNI intersubject correspondency, then the term “fine-grained” seems inappropriate. In either case, the question of the grain of the representational patterns should be explicitly discussed.
(2) What is the content of the event representations?
The Hidden Markov Model is great for capturing the boundaries between events. However, it does not capture the meaning and relationships between the event representations. It would be great to see the full time-by-time representational dissimilarity matrices (RDMs; or pattern similarity matrices) for multiple regions (and for single subjects and averaged across subjects). It would also be useful to average the dissimilarities within each pair of events to obtain event-by-event RDMs. These should reveal, when events recur in the movie, and the degree of similarity of different events in each brain region. If each event were unique in the movie experience, these RDMs would have a diagonal structure. Analysing the content of the event representations in some way seems essential to the interpretation that the patterns represent events.
(3) Why do the time-by-time pattern similarity matrices look so low-dimensional?
The pattern correlations shown in Figure 2 for precuneus and V1 are very high in absolute value and seem to span the entire range from -1 to 1. (Are the patterns averaged across all subjects?) It looks like two events either have highly correlated or highly anticorrelated patterns. This would suggest that there are only two event representations and each event falls into one of two categories. Perhaps there are intermediate values, but the structure of these matrices looks low-dimensional (essentially 1 dimensional) to me. The strong negative correlations might be produced by the way the data are processed, which could be more clearly described. For example, if the ensemble of patterns were centered in the response space by subtracting the mean pattern from each pattern, then strong negative correlations would arise.
I am wondering to what extent these matrices might reflect coarse-scale overall activation fluctuations rather than detailed representations of individual events. The correlation distance removes the mean from each pattern, but usually different voxels respond with different gains, so activation scales rather than translates the pattern up. When patterns are centered in response space, 1-dimensional overall activation dynamics can lead to the appearance of correlated and anticorrelated pattern states (along with intermediate correlations) as seen here.
This concern relates also to points (1) and (2) above and could be addressed by analysing fine-grained within-subject patterns and the content of the event representations.
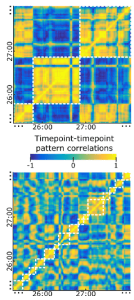
Detail from Figure 2: Time-by-time regional spatial-pattern correlation matrices.
Precuneus (top) and V1 (bottom).
(4) Do brain regions really represent a discrete sequence of events by a discrete sequence of patterns?
The paper currently claims to show that brain regions represent events as static patterns, with sudden switches at the event boundaries. However, this is not something that is demonstrated from the data, rather it is the assumption built into the Hidden Markov Model.
I very much like the Hidden Markov Model, because it provides a data-driven way to discover the event boundaries. The model assumption of static patterns and sudden switches are fine for this purpose because they may provide an approximation to what is really going on. Sudden switches are plausible, since transitions between events are sudden cognitive phenomena. However, it seems unlikely that patterns are static within events. This claim should be removed or substantiated by an inferential comparison of the static-pattern sequence model with an alternative model that allows for dynamic patterns within each event.
(5) Why use the contrast of within- and between-event pattern correlation in held-out subjects as the criterion for evaluating the performance of the Hidden Markov Model?
If patterns are assumed to be aligned between subjects, the Hidden Markov Model could be used to directly predict the pattern time course in a held-out subject. (Predicting the average of the training subjects’ pattern time courses would provide a noise ceiling.) The within- minus between-event pattern correlation has the advantage that it doesn’t require the assumption of intersubject pattern alignment, but this advantage appears not to be exploited here. The within- minus between-event pattern correlation seems problematic here because patterns acquired closer in time tend to be more similar (Henriksson et al. 2015). First, the average within-event correlation should always tend to be larger than the average between-event correlation (unless the average between-event correlation were estimated from the same distribution of temporal lags). Such a positive bias would be no problem for comparing between different segmentations. However, if temporally close patterns are more similar, then even in the absence of any event structure, we expect that a certain number of events best captures the similarity among temporally closeby patterns. The inference of the best number of events would then be biased toward the number of events, which best captures the continuous autocorrelation.
(6) More details on the recall reactivation
Fig. 5a is great. However, this is a complex analysis and it would be good to see this in all subjects and to also see the movie-to-recall pattern similarity matrix, with the human annotations-based and Hidden Markov Model-based time-warp trajectories superimposed. This would enable us to better understand the data and how the Hidden Markov Model arrives at the assignment of corresponding events.
In addition, it would be good to show statistically, that the Hidden Markov Model predicts the content correspondence between movie and recall representations consistently with the human annotations.
(7) fMRI is a hemodynamic measure, not “neural data”.
“Using a data-driven event segmentation model that can identify temporal structure directly from neural measurements”; “Our results are the first to demonstrate a number of key predictions of event segmentation theory (Zacks et al., 2007) directly from neural data”
There are a couple of other places, where “neural data” is used. Better terms include “fMRI data” and “brain activity patterns”.
(8) Is the structure of the multi-time-scale event segmentation a tree?
Do all regions that represent the same time-scale have the same event boundaries? Or do they provide alternative temporal segmentations? If it is the former, do short-time-scale regions strictly subdivide the segmentation of longer-time-scale regions, thus making the event structure a tree? Fig. 1 appears to be designed so as not to imply this claim. Data, of course, is noisy, so we don’t expect a perfect tree to emerge in the analysis, even if our brains did segment experience into a perfect tree. It would be good to perform an explicit statistical comparison between the temporal-tree event segmentation hypothesis and the more general multi-time-scale event segmentation hypothesis.
(9) Isn’t it a foregone conclusion that longer-time-scale regions’ temporal boundaries will match better to human annotated boundaries?
“We then measured, for each brain searchlight, what fraction of its neurally-defined boundaries were close to (within three time points of) a human-labeled event boundary.”
For a region with twice as many boundaries as another region, this fraction is halved even if both regions match all human labeled events. This analysis therefore appears strongly confounded by the number of events a regions represents.
The confound could be removed by having humans segment the movie at multiple scales (or having them segment at a short time scale and assign saliency ratings to the boundaries). The number of events could then be matched before comparing segmentations between human observers and brain regions.
Conversely, and without requiring more human annotations, the HMM could be constrained to the number of events labelled by humans for each searchlight location. This would ensure that the fraction of matches to human observers’ boundaries can be compared between regions.
(10) Hippocampus response does not appear to be “triggered” by the end of the event, but starts much earlier.
The hemodynamic peak is about 2-3 s after the event boundary, so we should expect the neural activity to begin well before the event boundary.
(11) Is the time scale a region represents reflected in the temporal power spectrum of spontaneous fluctuations?
The studies presenting such evidence are cited, but it would be good to look at the temporal power spectrum also for the present data and relate these two perspectives. I don’t think the case for event representation by static patterns is quite compelling (yet). Looking at the data also from this perspective may help us get a fuller picture.
(12) The title and some of the terminology is ambiguous
The title “Discovering event structure in continuous narrative perception and memory” is, perhaps intentionally, ambiguous. It is unclear who or what “discovers” the event structure. On the one hand, the brain that discovers event structure in the stream of experience. On the other hand, the Hidden Markov Model discovers good segmentations of regional pattern time courses. Although both interpretations work in retrospect, I would prefer a title that makes a point that’s clear from the beginning.
On a related note, the phrase “data-driven event segmentation model” suggests that the model performs the task of segmenting the sensory stream into events. This was initially confusing to me. In fact, what is used here is a brain-data-driven pattern time course segmentation model.
(13) Selection bias?
I was wondering about the possibility of selection bias (showing the data selected by brain mapping, which is biased by the selection process) for some of the figures, including Figs. 2, 4, and 7. It’s hard to resist illustrating the effects by showing selected data, but it can be misleading. Are the analyses for single searchlights? Could they be crossvalidated?
(14) Cubic searchlight
A spherical or surface-based searchlight would the better than a (2.1 cm)3 cube.
– Nikolaus Kriegeskorte
Acknowledgement
I thank Aya Ben-Yakov for discussing this paper with me.
One thought on “Discrete-event-sequence model reveals the multi-time-scale brain representation of experience and recall”